Molecular and physiological mechanisms of heat and drought stress tolerance in maize (Zea mays L.)
Keywords:
ABA signaling, Climate resilience, Drought stress, Heat stress, Maize (Zea mays L.), Stress toleranceAbstract
Maize (Zea mays L.) is one of the most important cereal crops worldwide, yet its growth and productivity are increasingly threatened by abiotic stresses, particularly drought and heat. These stresses, often occurring simultaneously under field conditions, severely impair photosynthesis, disrupt cellular homeostasis, and reduce yield potential. This review summarizes recent advances in understanding the physiological, biochemical, and molecular mechanisms underlying maize responses and tolerance to drought and heat stress. Key signaling pathways involving abscisic acid (ABA), calcium, reactive oxygen species (ROS), and mitogen-activated protein kinases (MAPKs) play central roles in stress perception and signal transduction. Transcription factors such as DREB, NAC, bZIP, HSF, and WRKY families coordinate downstream gene expression, regulating protective responses including osmotic adjustment, antioxidant defense, and protein stabilization. Moreover, the involvement of epigenetic modifications and transcriptional reprogramming highlights the complexity of maize adaptation and stress memory. Crosstalk between drought- and heat-responsive networks, exemplified by shared regulators such as ZmDREB2A, provides new insight into combined stress tolerance. Integration of multi-omics technologies, genome editing tools, and precision breeding holds great promise for developing climate-resilient maize varieties. Overall, a comprehensive understanding of these mechanisms is crucial for sustaining maize productivity and global food security under the escalating impacts of climate change.
References
1. Lamaoui M, Jemo M, Datla R, Bekkaoui F. 2018. Heat and drought stresses in crops and approaches for their mitigation. Frontiers in Chemistry 6:26
2. Ray D, Gerber J, MacDonald G, West P. 2015. Climate variation explains a third of global crop yield variability. Nat Commun 6: 5989.
3. Masson-Delmotte V, Zhai P, Pörtner H-O, Roberts D, Skea J, Shukla PR, Pirani A, Moufouma-Okia W, Péan C, Pidcock R. 2018. Global warming of 1.5 C. An IPCC Special Report on the impacts of global warming of 1:43-50
4. Yan J, Tan B-C. 2019. Maize biology: From functional genomics to breeding application. Journal of Integrative Plant Biology 61:654
5. Matthews HD, Wynes S. 2022. Current global efforts are insufficient to limit warming to 1.5 C. Science 376:1404-1409
6. Yang Z, Cao Y, Shi Y, Qin F, Jiang C, Yang S. 2023. Genetic and molecular exploration of maize environmental stress resilience: Toward sustainable agriculture. Molecular Plant 16:1496-1517
7. Farooqi MQU, Nawaz G, Wani SH, Choudhary JR, Rana M, Sah RP, Afzal M, Zahra Z, Ganie SA, Razzaq A. 2022. Recent developments in multi-omics and breeding strategies for abiotic stress tolerance in maize (Zea mays L.). Frontiers in Plant Science 13:965878
8. Zandalinas SI, Sengupta S, Fritschi FB, Azad RK, Nechushtai R, Mittler R. 2021. The impact of multifactorial stress combination on plant growth and survival. New Phytologist 230:1034-1048
9. Hasegawa PM, Bressan RA, Zhu J-K, Bohnert HJ. 2000. Plant cellular and molecular responses to high salinity. Annual Review of Plant Biology 51:463-499
10. Zhu J-K. 2016. Abiotic stress signaling and responses in plants. Cell 167:313-324
11. Ali M, Kaderbek T, Khan MA, Skalicky M, Brestic M, Elsabagh M, El Sabagh A. 2025. Biosynthesis and multifaceted roles of reactive species in plant defense mechanisms during environmental cues. Plant Stress:101102
12. Shabala S, Bose J, Fuglsang AT, Pottosin I. 2016. On a quest for stress tolerance genes: membrane transporters in sensing and adapting to hostile soils. Journal of Experimental Botany 67:1015-1031
13. Hatfield J, Prueger J. 2015. Temperature extremes: effect on plant growth and development. Weather Clim Extrem 10: 4-10.
14. Ding Y, Yang S. 2022. Surviving and thriving: How plants perceive and respond to temperature stress. Developmental Cell 57:947-958
15. Lobell DB, Deines JM, Tommaso SD. 2020. Changes in the drought sensitivity of US maize yields. Nature Food 1:729-735
16. Simpkins G. 2020. Maize sensitivity to drought. Nature Reviews Earth & Environment 1:625-625
17. Aslam M, Maqbool MA, Cengiz R. 2015. Drought stress in maize (Zea mays L.) Effects, resistance mechanisms, global achievements and. Cham: Springer
18. Saidi Y, Peter M, Finka A, Cicekli C, Vigh L, Goloubinoff P. 2010. Membrane lipid composition affects plant heat sensing and modulates Ca2+-dependent heat shock response. Plant signaling & behavior 5:1530-1533
19. Ma Y, Dai X, Xu Y, Luo W, Zheng X, Zeng D, Pan Y, Lin X, Liu H, Zhang D. 2015. COLD1 confers chilling tolerance in rice. Cell 160:1209-1221
20. Ding Y, Yang H, Wu S, Fu D, Li M, Gong Z, Yang S. 2022. CPK28-NLP7 module integrates cold-induced Ca2+ signal and transcriptional reprogramming in Arabidopsis. Science Advances 8:eabn7901
21. Ding Y, Shi Y, Yang S. 2020. Molecular regulation of plant responses to environmental temperatures. Molecular Plant 13:544-564
22. Kan Y, Lin H-X. 2021. Molecular regulation and genetic control of rice thermal response. The Crop Journal 9:497-505
23. Singh A, Pandey H, Pandey S, Lal D, Chauhan D, Aparna, Antre SH, B S, Kumar A. 2023. Drought stress in maize: stress perception to molecular response and strategies for its improvement. Functional & Integrative Genomics 23:296
24. Zhao C, Liu B, Piao S, Wang X, Lobell DB, Huang Y, Huang M, Yao Y, Bassu S, Ciais P. 2017. Temperature increase reduces global yields of major crops in four independent estimates. Proceedings of the National Academy of Sciences 114:9326-9331
25. Hatfield JL, Prueger JH. 2015. Temperature extremes: Effect on plant growth and development. Weather and climate extremes 10:4-10
26. Mittler R, Finka A, Goloubinoff P. 2012. How do plants feel the heat? Trends in Biochemical Sciences 37:118-125
27. Zhang H, Zhou J-F, Kan Y, Shan J-X, Ye W-W, Dong N-Q, Guo T, Xiang Y-H, Yang Y-B, Li Y-C. 2022. A genetic module at one locus in rice protects chloroplasts to enhance thermotolerance. Science 376:1293-1300
28. Kan Y, Mu X-R, Zhang H, Gao J, Shan J-X, Ye W-W, Lin H-X. 2022. TT2 controls rice thermotolerance through SCT1-dependent alteration of wax biosynthesis. Nature Plants 8:53-67
29. Zhao Y, Du H, Wang Y, Wang H, Yang S, Li C, Chen N, Yang H, Zhang Y, Zhu Y. 2021. The calcium‐dependent protein kinase ZmCDPK7 functions in heat‐stress tolerance in maize. Journal of Integrative Plant Biology 63:510-527
30. Ohama N, Sato H, Shinozaki K, Yamaguchi-Shinozaki K. 2017. Transcriptional regulatory network of plant heat stress response. Trends in Plant Science 22:53-65
31. Li B, Gao K, Ren H, Tang W. 2018. Molecular mechanisms governing plant responses to high temperatures. Journal of Integrative Plant Biology 60:757-779
32. Andrási N, Pettkó-Szandtner A, Szabados L. 2021. Diversity of plant heat shock factors: regulation, interactions, and functions. Journal of Experimental Botany 72:1558-1575
33. Liu HC, Liao HT, Charng YY. 2011. The role of class A1 heat shock factors (HSFA1s) in response to heat and other stresses in Arabidopsis. Plant, Cell & Environment 34:738-751
34. Lin YX, Jiang HY, Chu ZX, Tang XL, Zhu SW, Cheng BJ. 2011. Genome-wide identification, classification and analysis of heat shock transcription factor family in maize. BMC Genomics 12:76
35. Li H-c, Zhang H-n, Li G-l, Liu Z-h, Zhang Y-m, Zhang H-m, Guo X-l. 2015. Expression of maize heat shock transcription factor gene ZmHsf06 enhances the thermotolerance and drought-stress tolerance of transgenic Arabidopsis. Functional Plant Biology 42:1080-1091
36. Li G-l, Zhang H-n, Shao H, Wang G-y, Zhang Y-y, Zhang Y-j, Zhao L-n, Guo X-l, Sheteiwy MS. 2019. ZmHsf05, a new heat shock transcription factor from Zea mays L. improves thermotolerance in Arabidopsis thaliana and rescues thermotolerance defects of the athsfa2 mutant. Plant Science 283:375-384
37. Zhang H, Li G, Hu D, Zhang Y, Zhang Y, Shao H, Zhao L, Yang R, Guo X. 2020. Functional characterization of maize heat shock transcription factor gene ZmHsf01 in thermotolerance. PeerJ 8:e8926
38. Jacob P, Hirt H, Bendahmane A. 2017. The heat‐shock protein/chaperone network and multiple stress resistance. Plant Biotechnology Journal 15:405-414
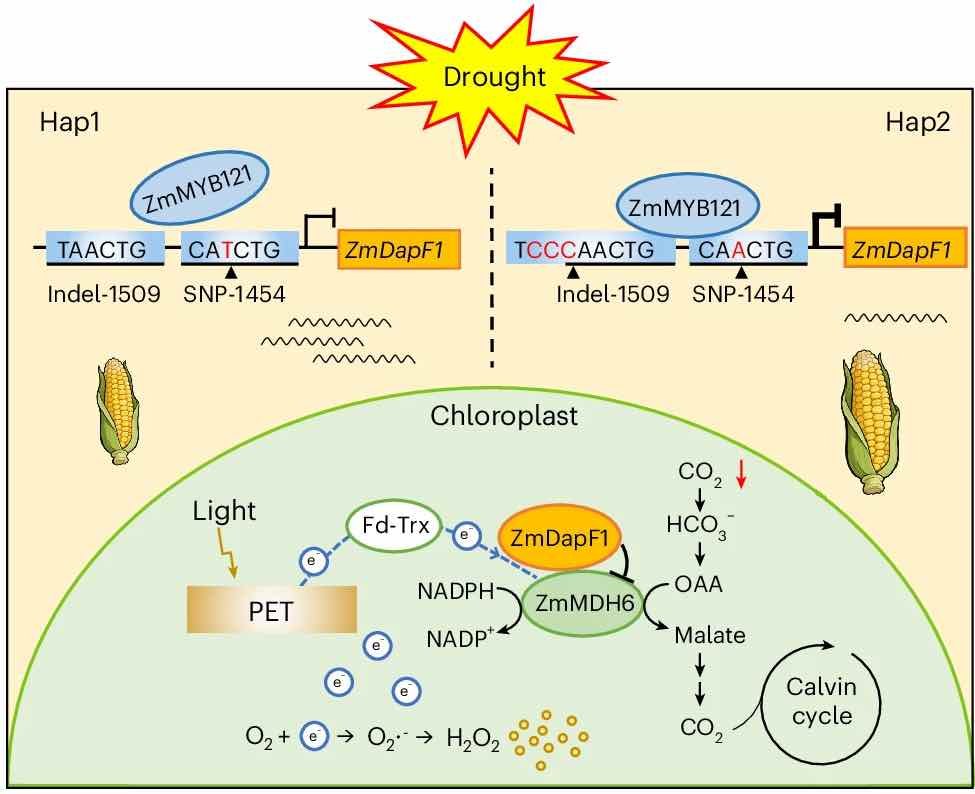
39. Lian Y, Yang S, Tian T, Yang Z, Liu S, Fu X, Liu C, Zhu T, Wang Y, Bai Y. 2025. Natural variation in ZmDapF1 enhances maize drought resilience. Nature Plants:1-14
40. Nieto-Sotelo J, Martínez LM, Ponce G, Cassab GI, Alagón A, Meeley RB, Ribaut J-M, Yang R. 2002. Maize HSP101 plays important roles in both induced and basal thermotolerance and primary root growth. The Plant Cell 14:1621-1633
41. Li M, Lin L, Zhang Y, Sui N. 2019. ZmMYB31, a R2R3-MYB transcription factor in maize, positively regulates the expression of CBF genes and enhances resistance to chilling and oxidative stress. Molecular Biology Reports 46:3937-3944
42. Li JY, Yang C, Xu J, Lu HP, Liu JX. 2023. The hot science in rice research: How rice plants cope with heat stress. Plant, Cell & Environment 46:1087-1103
43. Lia Z, Tangb J, Srivastavaa R, Basshamb DC, Howella SH. 2020. The transcription factor bZIP60 links the unfolded protein response (UPR) to the heat stress response (HSR) in maize. The Plant Cell 32:3559-75
44. Li Z, Srivastava R, Tang J, Zheng Z, Howell SH. 2018. Cis-effects condition the induction of a major unfolded protein response factor, ZmbZIP60, in response to heat stress in maize. Frontiers in Plant Science 9:833
45. Xie C, Yang L, Jia G, Yan K, Zhang S, Yang G, Wu C, Gai Y, Zheng C, Huang J. 2022. Maize HEAT UP-REGULATED GENE 1 plays vital roles in heat stress tolerance. Journal of Experimental Botany 73:6417-6433
46. Liu JX, Howell SH. 2016. Managing the protein folding demands in the endoplasmic reticulum of plants. New Phytologist 211:418-428
47. Sakuma Y, Maruyama K, Qin F, Osakabe Y, Shinozaki K, Yamaguchi-Shinozaki K. 2006. Dual function of an Arabidopsis transcription factor DREB2A in water-stress-responsive and heat-stress-responsive gene expression. Proceedings of the National Academy of Sciences 103:18822-18827
48. Qin F, Kakimoto M, Sakuma Y, Maruyama K, Osakabe Y, Tran LSP, Shinozaki K, Yamaguchi‐Shinozaki K. 2007. Regulation and functional analysis of ZmDREB2A in response to drought and heat stresses in Zea mays L. The Plant Journal 50:54-69
49. Gupta A, Rico-Medina A, Caño-Delgado AI. 2020. The physiology of plant responses to drought. Science 368:266-269
50. Agurla S, Gahir S, Munemasa S, Murata Y, Raghavendra AS. 2018. Mechanism of stomatal closure in plants exposed to drought and cold stress. Survival strategies in extreme cold and desiccation: adaptation mechanisms and their applications:215-232
51. Ma Y, Szostkiewicz I, Korte A, Moes D, Yang Y, Christmann A, Grill E. 2009. Regulators of PP2C phosphatase activity function as abscisic acid sensors. Science 324:1064-1068
52. Park S-Y, Fung P, Nishimura N, Jensen DR, Fujii H, Zhao Y, Lumba S, Santiago J, Rodrigues A, Chow T-fF. 2009. Abscisic acid inhibits type 2C protein phosphatases via the PYR/PYL family of START proteins. Science 324:1068-1071
53. Uno Y, Furihata T, Abe H, Yoshida R, Shinozaki K, Yamaguchi-Shinozaki K. 2000. Arabidopsis basic leucine zipper transcription factors involved in an abscisic acid-dependent signal transduction pathway under drought and high-salinity conditions. Proceedings of the National Academy of Sciences 97:11632-11637
54. Geiger D, Scherzer S, Mumm P, Stange A, Marten I, Bauer H, Ache P, Matschi S, Liese A, Al-Rasheid KA. 2009. Activity of guard cell anion channel SLAC1 is controlled by drought-stress signaling kinase-phosphatase pair. Proceedings of the National Academy of Sciences 106:21425-21430
55. Wang Y-G, Fu F-L, Yu H-Q, Hu T, Zhang Y-Y, Tao Y, Zhu J-K, Zhao Y, Li W-C. 2018. Interaction network of core ABA signaling components in maize. Plant Molecular Biology 96:245-263
56. Wu Q, Wang M, Shen J, Chen D, Zheng Y, Zhang W. 2019. ZmOST1 mediates abscisic acid regulation of guard cell ion channels and drought stress responses. Journal of Integrative Plant Biology 61:478-491
57. Xiang Y, Sun X, Gao S, Qin F, Dai M. 2017. Deletion of an endoplasmic reticulum stress response element in a ZmPP2C-A gene facilitates drought tolerance of maize seedlings. Molecular Plant 10:456-469
58. Li XD, Gao YQ, Wu WH, Chen LM, Wang Y. 2022. Two calcium‐dependent protein kinases enhance maize drought tolerance by activating anion channel ZmSLAC1 in guard cells. Plant Biotechnology Journal 20:143-157
59. Gao H, Cui J, Liu S, Wang S, Lian Y, Bai Y, Zhu T, Wu H, Wang Y, Yang S. 2022. Natural variations of ZmSRO1d modulate the trade-off between drought resistance and yield by affecting ZmRBOHC-mediated stomatal ROS production in maize. Molecular Plant 15:1558-1574
60. Soma F, Takahashi F, Suzuki T, Shinozaki K, Yamaguchi-Shinozaki K. 2020. Plant Raf-like kinases regulate the mRNA population upstream of ABA-unresponsive SnRK2 kinases under drought stress. Nature Communications 11:1373
61. Takahashi Y, Zhang J, Hsu P-K, Ceciliato PH, Zhang L, Dubeaux G, Munemasa S, Ge C, Zhao Y, Hauser F. 2020. MAP3Kinase-dependent SnRK2-kinase activation is required for abscisic acid signal transduction and rapid osmotic stress response. Nature Communications 11:12
62. Chen X, Ding Y, Yang Y, Song C, Wang B, Yang S, Guo Y, Gong Z. 2021. Protein kinases in plant responses to drought, salt, and cold stress. Journal of Integrative Plant Biology 63:53-78
63. Lu F, Wang K, Yan L, Peng Y, Qu J, Wu J, Cao Y, Yang Q, Fu F, Yu H. 2020. Isolation and characterization of maize ZmPP2C26 gene promoter in drought-response. Physiology and Molecular Biology of Plants 26:2189-2197
64. Lu F, Li W, Peng Y, Cao Y, Qu J, Sun F, Yang Q, Lu Y, Zhang X, Zheng L. 2022. ZmPP2C26 alternative splicing variants negatively regulate drought tolerance in maize. Frontiers in Plant Science 13:851531
65. Guo Y, Shi Y, Wang Y, Liu F, Li Z, Qi J, Wang Y, Zhang J, Yang S, Wang Y. 2023. The clade F PP2C phosphatase ZmPP84 negatively regulates drought tolerance by repressing stomatal closure in maize. New Phytologist 237:1728-1744
66. Guo M, Rupe MA, Wei J, Winkler C, Goncalves-Butruille M, Weers BP, Cerwick SF, Dieter JA, Duncan KE, Howard RJ. 2014. Maize ARGOS1 (ZAR1) transgenic alleles increase hybrid maize yield. Journal of Experimental Botany 65:249-260
67. Shi J, Habben JE, Archibald RL, Drummond BJ, Chamberlin MA, Williams RW, Lafitte HR, Weers BP. 2015. Overexpression of ARGOS genes modifies plant sensitivity to ethylene, leading to improved drought tolerance in both Arabidopsis and maize. Plant Physiology 169:266-282
68. Shi J, Gao H, Wang H, Lafitte HR, Archibald RL, Yang M, Hakimi SM, Mo H, Habben JE. 2017. ARGOS 8 variants generated by CRISPR‐Cas9 improve maize grain yield under field drought stress conditions. Plant Biotechnology Journal 15:207-216
69. Habben JE, Bao X, Bate NJ, DeBruin JL, Dolan D, Hasegawa D, Helentjaris TG, Lafitte RH, Lovan N, Mo H. 2014. Transgenic alteration of ethylene biosynthesis increases grain yield in maize under field drought‐stress conditions. Plant biotechnology journal 12:685-693
70. Seeve CM, Cho IJ, Hearne LB, Srivastava GP, Joshi T, Smith DO, Sharp RE, Oliver MJ. 2017. Water‐deficit‐induced changes in transcription factor expression in maize seedlings. Plant, Cell & Environment 40:686-701
71. Leng P, Zhao J. 2020. Transcription factors as molecular switches to regulate drought adaptation in maize. Theoretical and Applied Genetics 133:1455-1465
72. Ma H, Liu C, Li Z, Ran Q, Xie G, Wang B, Fang S, Chu J, Zhang J. 2018. ZmbZIP4 contributes to stress resistance in maize by regulating ABA synthesis and root development. Plant Physiology 178:753-770
73. Li Z, Liu C, Zhang Y, Wang B, Ran Q, Zhang J. 2019. The bHLH family member ZmPTF1 regulates drought tolerance in maize by promoting root development and abscisic acid synthesis. Journal of Experimental Botany 70:5471-5486
74. Liu S, Wang X, Wang H, Xin H, Yang X, Yan J, Li J, Tran L-SP, Shinozaki K, Yamaguchi-Shinozaki K. 2013. Genome-wide analysis of ZmDREB genes and their association with natural variation in drought tolerance at seedling stage of Zea mays L. PLoS Genetics 9:e1003790
75. Mao H, Wang H, Liu S, Li Z, Yang X, Yan J, Li J, Tran L-SP, Qin F. 2015. A transposable element in a NAC gene is associated with drought tolerance in maize seedlings. Nature Communications 6:8326
76. Xiang Y, Sun X, Bian X, Wei T, Han T, Yan J, Zhang A. 2021. The transcription factor ZmNAC49 reduces stomatal density and improves drought tolerance in maize. Journal of Experimental Botany 72:1399-1410
77. Tian T, Wang S, Yang S, Yang Z, Liu S, Wang Y, Gao H, Zhang S, Yang X, Jiang C. 2023. Genome assembly and genetic dissection of a prominent drought-resistant maize germplasm. Nature Genetics 55:496-506
78. Yang Y, Shi J, Chen L, Xiao W, Yu J. 2022. ZmEREB46, a maize ortholog of Arabidopsis WAX INDUCER1/SHINE1, is involved in the biosynthesis of leaf epicuticular very-long-chain waxes and drought tolerance. Plant Science 321:111256
79. Wang Z, Zhao X, Ren Z, Abou‐Elwafa SF, Pu X, Zhu Y, Dou D, Su H, Cheng H, Liu Z. 2022. ZmERF21 directly regulates hormone signaling and stress‐responsive gene expression to influence drought tolerance in maize seedlings. Plant, Cell & Environment 45:312-328
80. Castorina G, Domergue F, Chiara M, Zilio M, Persico M, Ricciardi V, Horner DS, Consonni G. 2020. Drought-responsive ZmFDL1/MYB94 regulates cuticle biosynthesis and cuticle-dependent leaf permeability. Plant Physiology 184:266-282
81. Feng X, Xiong J, Zhang W, Guan H, Zheng D, Xiong H, Jia L, Hu Y, Zhou H, Wen Y. 2022. ZmLBD5, a class‐II LBD gene, negatively regulates drought tolerance by impairing abscisic acid synthesis. The Plant Journal 112:1364-1376
82. Nelson DE, Repetti PP, Adams TR, Creelman RA, Wu J, Warner DC, Anstrom DC, Bensen RJ, Castiglioni PP, Donnarummo MG. 2007. Plant nuclear factor Y (NF-Y) B subunits confer drought tolerance and lead to improved corn yields on water-limited acres. Proceedings of the National Academy of Sciences 104:16450-16455
83. Su H, Cao Y, Ku L, Yao W, Cao Y, Ren Z, Dou D, Wang H, Ren Z, Liu H. 2018. Dual functions of ZmNF-YA3 in photoperiod-dependent flowering and abiotic stress responses in maize. Journal of Experimental Botany 69:5177-5189
84. Bruce WB, Edmeades GO, Barker TC. 2002. Molecular and physiological approaches to maize improvement for drought tolerance. Journal of Experimental Botany 53:13-25
85. Liu Q, Ding Y, Shi Y, Ma L, Wang Y, Song C, Wilkins KA, Davies JM, Knight H, Knight MR. 2021. The calcium transporter ANNEXIN1 mediates cold‐induced calcium signaling and freezing tolerance in plants. The EMBO Journal 40:e104559
86. Chang YN, Zhu C, Jiang J, Zhang H, Zhu JK, Duan CG. 2020. Epigenetic regulation in plant abiotic stress responses. Journal of Integrative Plant Biology 62:563-580
87. Forestan C, Farinati S, Zambelli F, Pavesi G, Rossi V, Varotto S. 2020. Epigenetic signatures of stress adaptation and flowering regulation in response to extended drought and recovery in Zea mays. Plant, Cell & Environment 43:55-75
88. Shang JY, He XJ. 2022. Chromatin‐remodeling complexes: Conserved and plant‐specific subunits in Arabidopsis. Journal of Integrative Plant Biology 64:499-515
89. Yu X, Meng X, Liu Y, Li N, Zhang A, Wang T-J, Jiang L, Pang J, Zhao X, Qi X. 2018. The chromatin remodeler ZmCHB101 impacts expression of osmotic stress-responsive genes in maize. Plant Molecular Biology 97:451-465
Downloads
Published
Data Availability Statement
The authors have nothing to report.
Issue
Section
License
Copyright (c) 2026 by the author(s)

This work is licensed under a Creative Commons Attribution 4.0 International License.
