Genome-wide identification and expression profiling of TALE gene family in banana under abiotic stresses and Foc 4 infection
Keywords:
Banana, Fusarium oxysporum, Foc 4, Genome-wide, MaTALE1, PEGAbstract
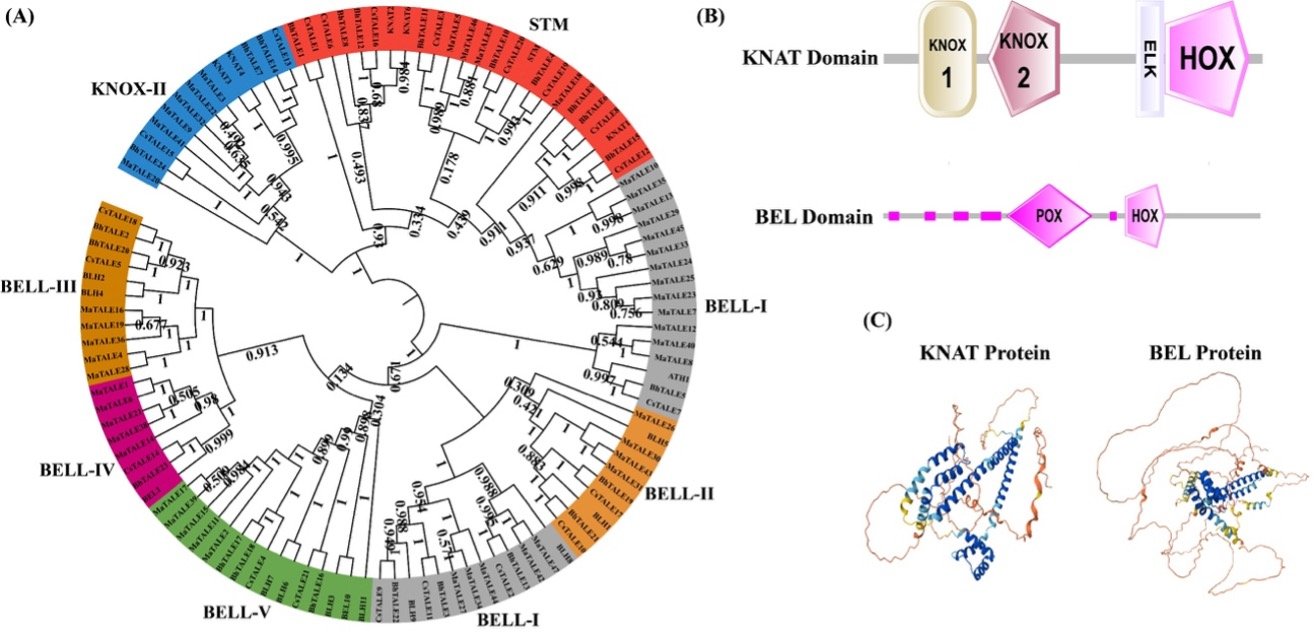
Three Amino Acid Loop Extension (TALE) is a member of the homeobox gene group. In addition to being crucial for growth and development, the TALE gene family is also critical for controlling how plants react to environmental stressors. Here, we used the banana genome database to extract 47 MaTALE genes. The evolutionary analysis grouped the sequences into seven subclades, named STM, KNOX-II, and BELL-I to BELL-V. Analysis of MaTALE promoters revealed that their cis-acting elements are associated primarily with hormone (especially ABA) and stress responsiveness. The STRING database further suggested that MaTALE proteins could work in tandem with other important TFs that play a significant role in plant growth. The expression analysis under drought, cold, and Foc 4 revealed their potential involvement in stress response. Exogenous ABA treatment significantly induced the expression of MaTALE1/14/20 genes. The transient expression assay showed that MaTALE1 resides in the nucleus. Our data offer a valuable resource for developing integrated management strategies against abiotic (drought, cold) and biotic (Foc 4) stresses affecting bananas in Southern China.
References
1. Bürglin TR. 1997. Analysis of TALE superclass homeobox genes (MEIS, PBC, KNOX, Iroquois, TGIF) reveals a novel domain conserved between plants and animals. Nucleic Acids Research 25:4173-4180
2. Gehring WJ. 1987. Homeo boxes in the study of development. Science 236:1245-1252
3. Qian B, Wang Q, Zhang C, Guo J, Yu Z, Han J, Xia H, Zhao R, Yin Y. 2024. Exploring the roles of TALE gene family in maize drought stress responses. Agronomy 14:1267
4. Jia P, Wang Y, Sharif R, Dong Q-l, Liu Y, Luan H-a, Zhang X-m, Guo S-p, Qi G-h. 2023. KNOTTED1-like homeobox (KNOX) transcription factors - hubs in a plethora of networks: A review. International Journal of Biological Macromolecules 253:126878
5. Jia P, Sharif R, Li Y, Sun T, Li S, Zhang X, Dong Q, Luan H, Guo S, Ren X, Qi G. 2023. The BELL1-like homeobox gene MdBLH14 from apple controls flowering and plant height via repression of MdGA20ox3. International Journal of Biological Macromolecules:124790
6. Scofield S, Dewitte W, Nieuwland J, Murray JAH. 2013. The Arabidopsis homeobox gene has cellular and meristem-organisational roles with differential requirements for cytokinin and CYCD3 activity. The Plant Journal 75:53-66
7. Ahmad S, Khan K, Saleh IA, Okla MK, Alaraidh IA, AbdElgawad H, Naeem M, Ahmad N, Fahad S. 2024. TALE gene family: identification, evolutionary and expression analysis under various exogenous hormones and waterlogging stress in Cucumis sativus L. BMC Plant Biology 24:564
8. Guo C, Quan S, Zhang Z, Kang C, Liu J, Niu J. 2022. Genome-wide identification, characterization and expression profile of TALE gene family in (Juglans regia L.). Scientia Horticulturae 297:110945
9. Testone G, Condello E, Verde I, Nicolodi C, Caboni E, Dettori MT, Vendramin E, Bruno L, Bitonti MB, Mele G. 2012. The peach (Prunus persica L. Batsch) genome harbours 10 KNOX genes, which are differentially expressed in stem development, and the class 1 KNOPE1 regulates elongation and lignification during primary growth. Journal of Experimental Botany 63:5417-5435
10. Wang S, Yamaguchi M, Grienenberger E, Martone PT, Samuels AL, Mansfield SD. 2020. The Class II KNOX genes KNAT3 and KNAT7 work cooperatively to influence deposition of secondary cell walls that provide mechanical support to Arabidopsis stems. The Plant Journal 101:293-309
11. Qin W, Yin Q, Chen J, Zhao X, Yue F, He J, Yang L, Liu L, Zeng Q, Lu F. 2020. The class II KNOX transcription factors KNAT3 and KNAT7 synergistically regulate monolignol biosynthesis in Arabidopsis. Journal of Experimental Botany 71:5469-5483
12. He JB, Zhao XH, Du PZ, Zeng W, Beahan CT, Wang YQ, Li HL, Bacic A, Wu AM. 2018. KNAT7 positively regulates xylan biosynthesis by directly activating IRX9 expression in Arabidopsis. Journal of Integrative Plant Biology 60:514-528
13. Sharif R, Su L, Chen X, Qi X. 2022. Involvement of auxin in growth and stress response of cucumber. Vegetable Research:1-9
14. Iannelli MA, Nicolodi C, Coraggio I, Fabriani M, Baldoni E, Frugis G. 2023. A Novel Role of Medicago truncatula KNAT3/4/5-like Class 2 KNOX Transcription Factors in Drought Stress Tolerance. International Journal of Molecular Sciences 24:12668
15. Zhang J-B, Wang Y, Zhang S-P, Cheng F, Zheng Y, Li Y, Li X-B. 2024. The BEL1-like transcription factor GhBLH5-A05 participates in cotton response to drought stress. The Crop Journal 12:177-187
16. Sun R, Qin T, Wall SB, Wang Y, Guo X, Sun J, Liu Y, Wang Q, Zhang B. 2023. Genome-wide identification of KNOX transcription factors in cotton and the role of GhKNOX4-A and GhKNOX22-D in response to salt and drought stress. International Journal of Biological Macromolecules 226:1248-1260
17. Li Y, Hu Y, Xiao W, Yan L, Feng J, Cao M, Si Y, Lyu J, Zhao Y, Li K, Wei Y, Hu H, Li W, Lü P, Wang W, Han Z, Xie J. 2026. A MaERF110-MaMYB308 transcriptional module negatively regulates lignin-mediated defence against Fusarium wilt in banana. Plant Biotechnology Journal n/a
18. Gulzar S, Xu Y, Fei FF, Yi Z, Huang C, Sharif R, Long Z, Lu Y, Zhan H, Xu C. 2026. Burkholderia seminalis suppresses Fusarium wilt infection in banana by modulating cell wall integrity. Pesticide Biochemistry and Physiology 218:106950
19. Zhang Y, Liu S, Mostert D, Yu H, Zhuo M, Li G, Zuo C, Haridas S, Webster K, Li M, Grigoriev IV, Yi G, Viljoen A, Li C, Ma L-J. 2024. Virulence of banana wilt-causing fungal pathogen Fusarium oxysporum tropical race 4 is mediated by nitric oxide biosynthesis and accessory genes. Nature Microbiology 9:2232-2243
20. Paul JY, Khanna H, Kleidon J, Hoang P, Geijskes J, Daniells J, Zaplin E, Rosenberg Y, James A, Mlalazi B. 2017. Golden bananas in the field: elevated fruit pro‐vitamin A from the expression of a single banana transgene. Plant Biotechnology Journal 15:520-532
21. Song Z, Lai X, Chen H, Wang L, Pang X, Hao Y, Lu W, Chen W, Zhu X, Li X. 2022. Role of MaABI5-like in abscisic acid-induced cold tolerance of ‘Fenjiao’ banana fruit. Horticulture Research 9: uhac130
22. Ji T, Li X, Meng G, Gu Y, Zhang Q, Liu L, Wu H, Yao Z, Zhang S, Wang Y, Zhang T, Wang X, Cao X, Li H, Liu Y, Wang X, Wang X, Sun S, Zhou M, Jia Q, Song K, Sun Z, Wu X-H, Niu K. 2020. The association between banana consumption and the depressive symptoms in Chinese general adult population: A cross-sectional study. Journal of Affective Disorders 264:1-6
23. Fan H, Dong H, Xu C, Liu J, Hu B, Ye J, Mai G, Li H. 2017. Pectin methylesterases contribute the pathogenic differences between races 1 and 4 of Fusarium oxysporum f. sp. cubense. Scientific Reports 7:13140
24. Wang J, Zhao P, Cheng B, Zhang Y, Shen Y, Wang X, Zhang Q, Lou Q, Zhang S, Wang B, Qi S, Li Y, Islam MM, Muhammad T, Zhang F, Liang Y. 2022. Identification of TALE transcription factor family and expression patterns related to fruit chloroplast development in tomato (Solanum lycopersicum L.). International Journal of Molecular Sciences 23:4507
25. Hussain S, Chang J, Li J, Chen X, Xie D, Zhang B. 2024. Transcriptome wide identification and expression analysis revealed BhTALE gene family regulates wax gourd (Benincasa hispida) response to low calcium and magnesium stress. Horticulturae 10:1083
26. Han Y, Zhang L, Yan L, Xiong X, Wang W, Zhang X-H, Min D-H. 2022. Genome-wide analysis of TALE superfamily in Triticum aestivum reveals TaKNOX11-A is involved in abiotic stress response. BMC Genomics 23:89
27. Ullah U, Shalmani A, Ilyas M, Raza A, Ahmad S, Shah AZ, Khan FU, AzizUd D, Bibi A, Rehman SU, Abbas Z, Buttar ZA. 2022. BZR proteins: identification, evolutionary and expression analysis under various exogenous growth regulators in plants. Molecular Biology Reports 49:12039-12053
28. Fei L, Liu J, Liao Y, Sharif R, Liu F, Lei J, Chen G, Zhu Z, Chen C. 2024. The CaABCG14 transporter gene regulates the capsaicin accumulation in Pepper septum. International Journal of Biological Macromolecules 280:136122
29. Zhao K, Zhang X, Cheng Z, Yao W, Li R, Jiang T, Zhou B. 2019. Comprehensive analysis of the three-amino-acid-loop-extension gene family and its tissue-differential expression in response to salt stress in poplar. Plant Physiology and Biochemistry 136:1-12
30. Wang Y, Zhao Y, Yan M, Zhao H, Zhang X, Yuan Z. 2020. Genome-wide identification and expression analysis of TALE gene family in pomegranate (Punica granatum L.). Agronomy 10:829
31. Zareen S, Ali A, Yun D-J. 2024. Significance of ABA biosynthesis in plant adaptation to drought stress. Journal of Plant Biology 67:175-184
32. Kim K, Lee J, Kim B, Shin J, Kang T-A, Kim W-C. 2022. GATA25, a novel regulator, accelerates the flowering time of Arabidopsis thaliana. Applied Biological Chemistry 65:28
33. Dai H, Zheng S, Zhang C, Huang R, Yuan L, Tong H. 2023. Identification and expression analysis of the KNOX genes during organogenesis and stress responseness in Camellia sinensis (L.) O. Kuntze. Molecular Genetics and Genomics 298:1559-1578
34. Li G, Manzoor MA, Wang G, Chen C, Song C. 2023. Comparative analysis of KNOX genes and their expression patterns under various treatments in Dendrobium huoshanense. Frontiers in Plant Science 14:1258533
35. Song X, Zhao Y, Wang J, Lu M-Z. 2021. The transcription factor KNAT2/6b mediates changes in plant architecture in response to drought via down-regulating GA20ox1 in Populus alba× P. glandulosa. Journal of Experimental Botany 72:5625-5637
36. Wang L, Yang X, Gao Y, Yang S. 2021. Genome-wide identification and characterization of TALE superfamily genes in soybean (Glycine max L.). International Journal of Molecular Sciences 22:4117
37. Razzaq A, Ashraf J, Malik W, Shaban M, Zhang R, Liang C-z, Hanif M, Abid MA, Qayyum A. 2020. In silico analyses of TALE transcription factors revealed its potential role for organ development and abiotic stress tolerance in cotton. 2020:1083-1093.
38. Zhang Q, Ahmad N, Li Z, He J, Wang N, Naeem M, Jin L, Yao N, Liu X. 2023. CtCYP71A1 promotes drought stress tolerance and lignin accumulation in safflower and Arabidopsis. Environmental and Experimental Botany 213:105430
39. Chen S, Jia Y, Yang Y, Liu H, Chen H, Liu J, Yin H, Zhuo R, Han X. 2025. Genome-wide analysis of the TsBLH gene family reveals TsBLH4 involved the regulation of abiotic stresses by interacting with KNOX6 in Toona sinensis. Plant Stress 15:100721
40. Zou M, Guo M, Zhou Z, Wang B, Pan Q, Li J, Zhou J-M, Li J. 2021. MPK3- and MPK6-mediated VLN3 phosphorylation regulates actin dynamics during stomatal immunity in Arabidopsis. Nature Communications 12:6474
41. Wei Y-S, Javed T, Liu T-T, Ali A, Gao S-J. 2025. Mechanisms of abscisic acid (ABA)-mediated plant defense responses: An updated review. Plant Stress 15:100724
42. Rathour M, Shumayla, Alok A, Upadhyay SK. 2022. Investigation of roles of TaTALE genes during development and stress response in bread wheat. Plants (Basel) 11:587
43. Liu P, Tang J, Lei Y, Zhang L, Ye J, Wang C, Zhou L, Liu Y, Wang Z, Jiang J, Chen F, Song A. 2025. Construction of the KNOX-BELL interaction network and functional analysis of CmBLH2 under cold stress in Chrysanthemum morifolium. International Journal of Biological Macromolecules 293:139365
44. Chen C, Wu Y, Li J, Wang X, Zeng Z, Xu J, Liu Y, Feng J, Chen H, He Y, Xia R. 2023. TBtools-II: A “one for all, all for one” bioinformatics platform for biological big-data mining. Molecular Plant 16:1733-1742
45. Sheraz A, Zhu H, Dong Q, Wang T, Zong S, Wang H, Ge L, Wu T. 2023. The superoxide dismutase (SOD) genes family mediates the response of Nilaparvata lugens to jinggangmycin and sugar. Frontiers in Physiology 14:1197395
46. Kelley LA, Mezulis S, Yates CM, Wass MN, Sternberg MJE. 2015. The Phyre2 web portal for protein modeling, prediction and analysis. Nature Protocols 10:845-858
47. Li F, Zhang L, Ji H, Xu Z, Zhou Y, Yang S. 2020. The specific W-boxes of GAPC5 promoter bound by TaWRKY are involved in drought stress response in wheat. Plant Science 296:110460
Downloads
Additional Files
Published
Data Availability Statement
All the data generated or analyzed during this study are included in this published article and its supplementary information file.
Issue
Section
License
Copyright (c) 2026 Rahat Sharif, Hangbo Cao, Huimin Song, Shazma Gulzar

This work is licensed under a Creative Commons Attribution 4.0 International License.
